
 |
Running HKL2000 at SSRLThe HKL2000 package brings together Denzo, Xdisplayf, and Scalepack using a graphical user interface. The information provided in this site is specific for running HKL2000 at SSRL. The HKL2000 on-line manual is available at the HKL Research Inc.. A pdf version of the HKL manual is also available. Experimenters that perform non-commercial research may freely use the program for data evaluation and processing, while collecting data at the beamline. These experimenters can also obtain a temporary, free of charge, license to (re)process data collected at synchrotron beamlines. This license can be obtained by contacting HKL Research, Inc. Experimenters from commercial and for-profit institutions, or researchers that would like to use the program for commercial research must have an appropriate commercial HKL license. NOTE: As of June 1, 2019, HKL3000 is the default version of HKL software available on the SSRL processing cluster. HKL2000 is still supported! To switch from HKL3000 to HKL2000, log into the data processing environment and type module switch -f HKL3000 HKL2000
Getting StartedHKL2000 program can be started from the data processing computers by typing HKL2000. This will launch the HKL2000 window and it will also show information about the other programs running on the machine in a red box as shown below. This box notifies the user about the status of the heavily used processors only and it can be usually ignored. 
Clicking Ok on this window gives you access to the HKL2000 launch window below. This window shows names of different detectors at SSRL and the small bar on the right can be moved up and down to see all the detectors. The default files for each of these detectors are available in these areas and they can be accessed by selecting the corresponding detector and clicking on Ok. Selection of a Q315 detector is shown on the window on the right side. If you need a copy of the default file, for this or any other of the detectors used at SSRL, please contact Irimpan Mathews.
This will bring the "HKL Main" window as shown below. This window gives access to different programs (Indexing, Integration. etc.) of the HKL2000 package from the menu bar on the top. If you are starting the HKL2000 program using a remote desktop it is difficult to see the whole window. To move the window up and down move the cursor to the left or right edge of the HKL2000 window and keep the left mouse button pressed while moving the window. Then, click the left mouse button again to keep the window in that view. 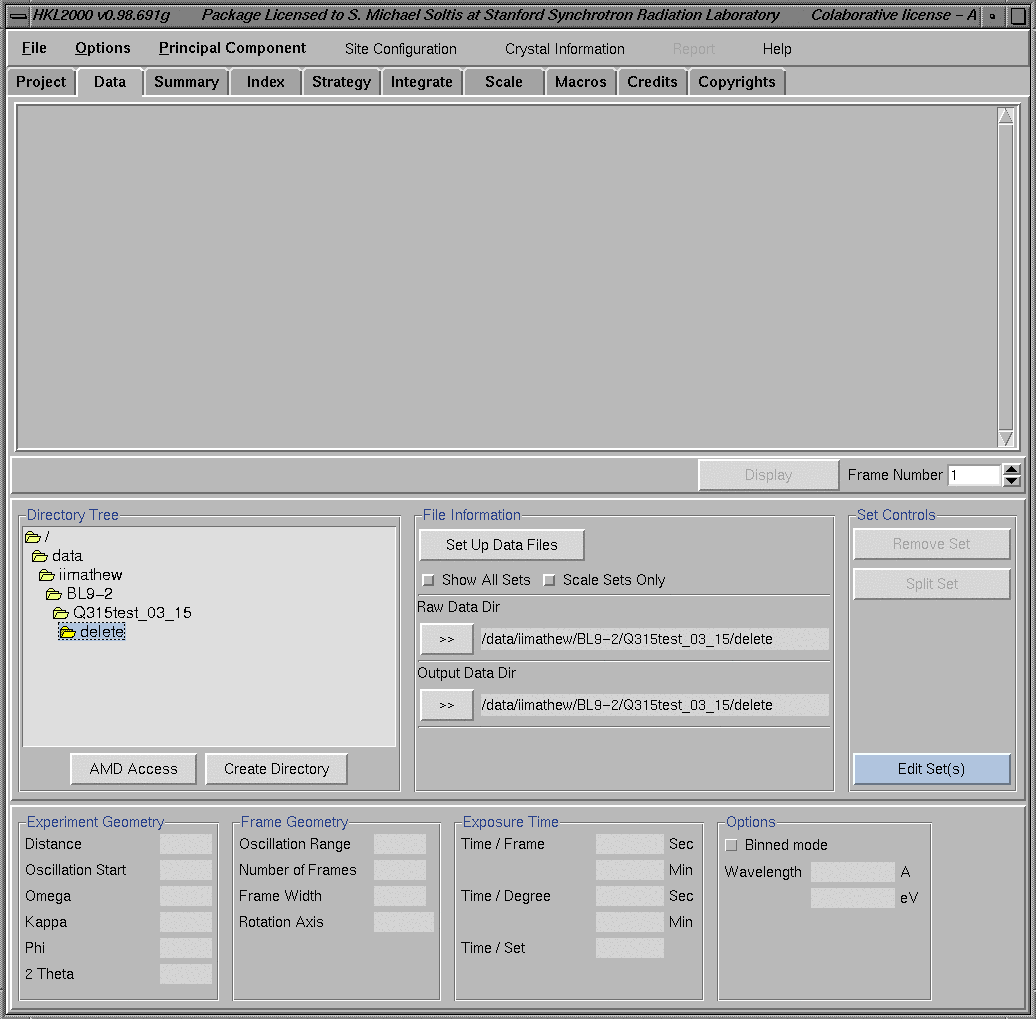
Clicking on "Help" will bring up a small window below with some information about data processing with HKL2000 program. If the detector is offset, the position of the direct beam may be typed in the "Site Configuration" window (select the "Site Configuration" menu.) 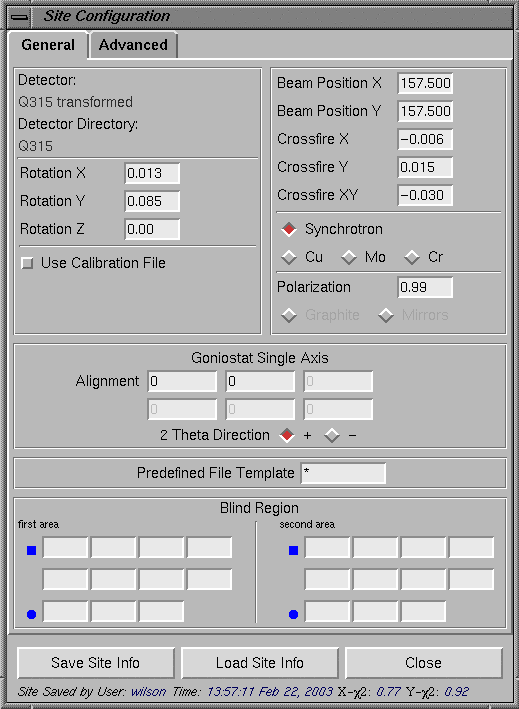
Close the configuration window by clicking on the "Close" button. This will save the newly typed information for the current session of the program. Please note that this file can/need not be saved using the "Save Site Info" button. Setting Data FilesUsually the HKL200 program is started either from the data collection directory or from a "process" sub directory under the data collection directory. For example, in the "HKL window" above, HKL2000 program is started from "/data/iimathew/BL9-2/Q315test_03_15/delete" directory. A "Directory Tree" is also displayed on the left side of the "HKL window". Set the "Raw Data Dir" to the directory where the images are located. The output files from HKL2000 processing will be directed to the "Output Data Dir". After this, click the Set Up Data Files button and program will bring all the image files in the "Raw Data Dir" as shown below. Sets of images for further processing can be selected or deselected by clicking on the corresponding sets. The display below shows the selection of 90 images corresponding to the "myoWI3_1_###.img [1-320]" data set. 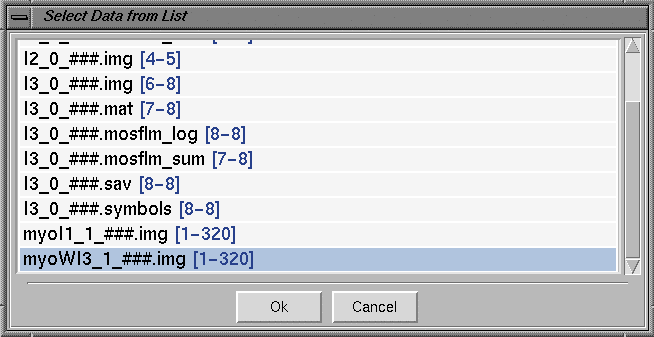
Clicking the "Ok" button will start a new window, as shown below (part of the window showing the relevant information is shown), giving details of the selected image set. If you selected 2 or 3 data sets (e.g., data collected at different energies for MAD experiments, low and high resolution passes) during the image selection process, you will see one window for each of the data sets. Sets of data from these windows for further processing can be selected or deselected by clicking on the Select button. A selected set can be removed by clicking on the "Remove Set" button on the lower portion of the "Main window". You can also choose data sets for either Denzo and/or Scale by selecting boxes on the left side of the Denzo and/or Scale names. For more information on processing several data sets
simultaneously, consult the HKL2000 manual.
During the above selection process, HKL2000 will also read all the relevant parameters from the image header and they will be displayed on the lower section of the general window (see the figure below). If the images were collected in the "binned" mode, the binned mode button in the "Other Options" section should be selected. 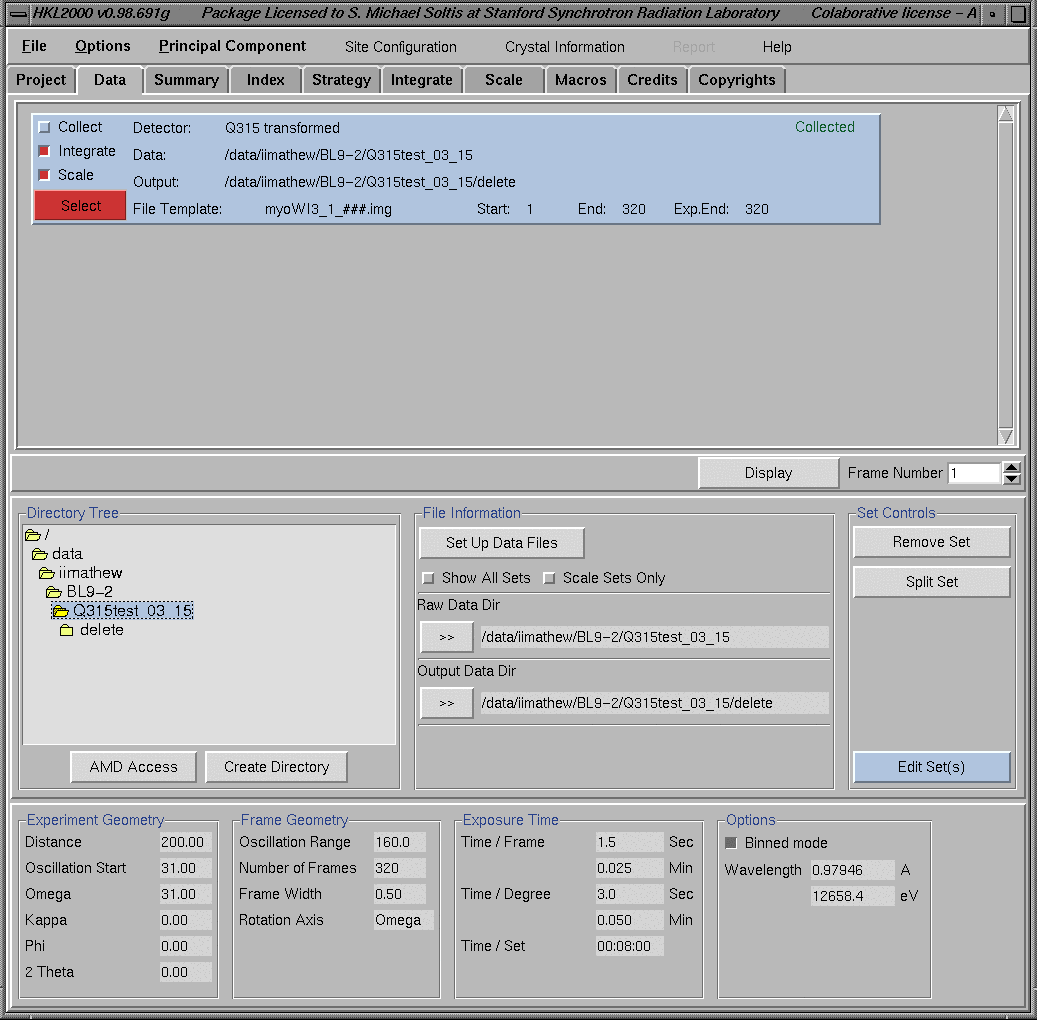
Clicking on the "Summary" tab shows a short summary of these parameters. The summary for the "myoWI3_1_###.img" images is shown below. (Note: If some of the parameters are incorrect, it can be corrected by typing their values in the above "Main HKL window" by clicking on the "Edit Set(S)" button. This is very useful for changing the last image number for an ongoing data collection. Please note that the current version of HKL200 doesn't work well if you change the image number after indexing the frames. Therefore, first use "Edit Set(S)" to correct the last image number for the current data collection and then index the images.) We can switch the displayed parameters "on" and "off" by clicking on the small box on the left hand side of their names in the summary control widget on the bottom part of the summary display.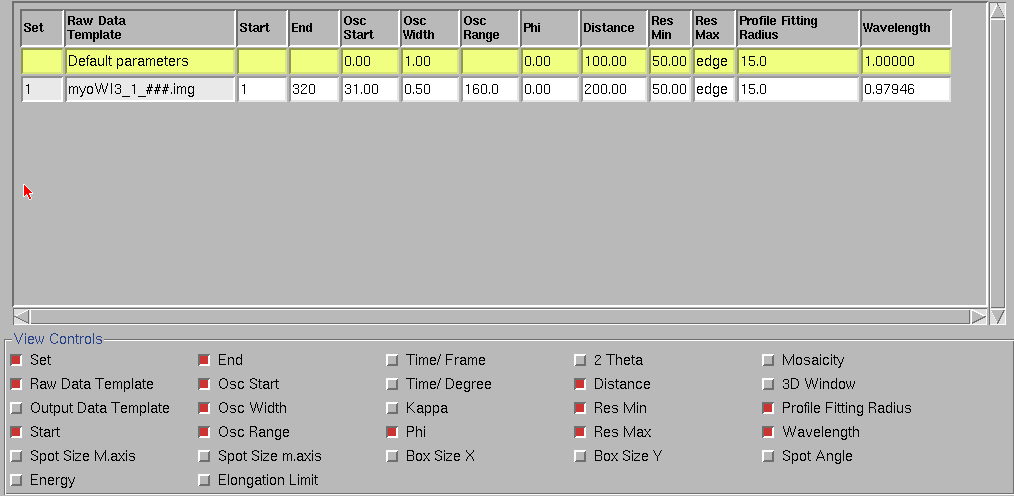
IndexingFirst we have to display an image and perform a "peak search" operation. We have to be in the "Main" menu (this menu can be accessed by clicking the "Menu" box on the left side of the "Summary" box) for displaying the necessary image file for peak search. After selecting the image click the Display button and this will launch the "xdisplayf" window (shown below). The peaks can be searched from the "xdisplayf" window. (Useful Tip: Peaks from a number of images for indexing can be selected by clicking on the "Frame" box of the xdisplayf window using the center mouse button before clicking "OK" for the peak search. This will show the next frame and the program automatically performs the "peak search" on this frame. Clicking "OK" at this point will write peaks from both images to the peaks file.) 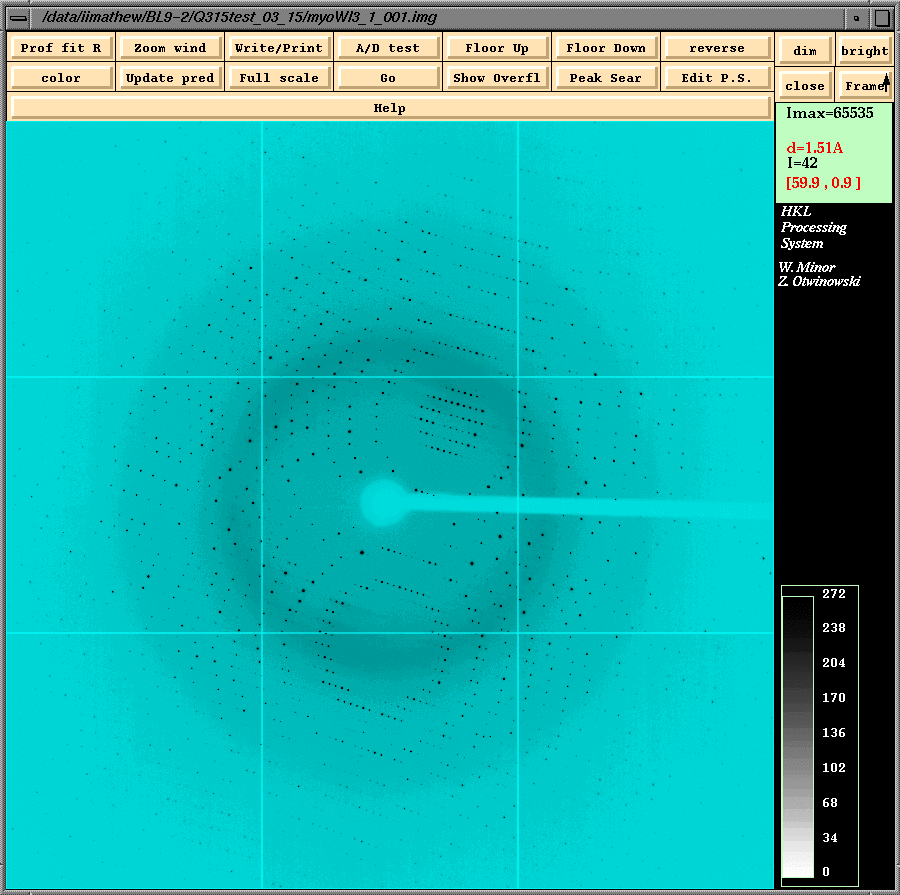
Selection of Index/Refinement from the "Main HKL2000 Window" will bring the following "Indexing/Refinement" window. 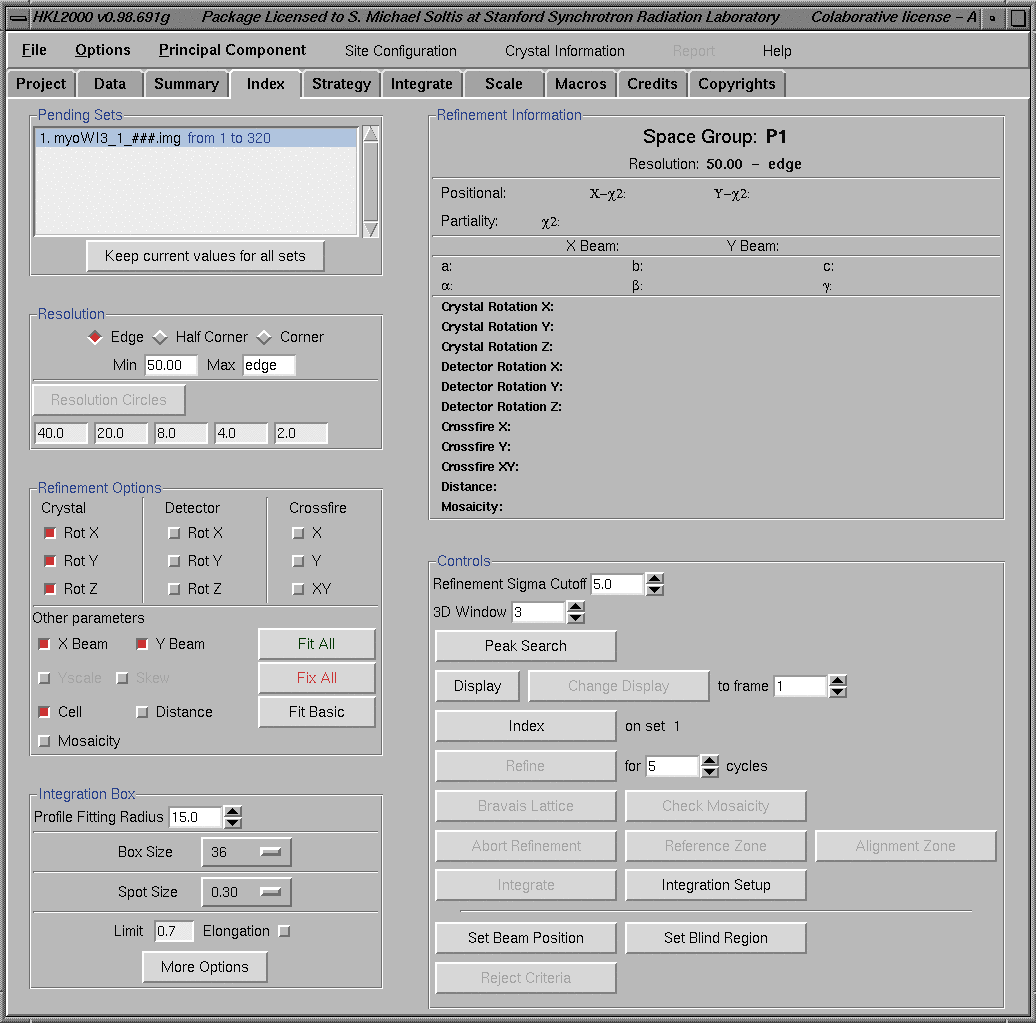
Click on the Index button to start the indexing process. Indexing process will bring a "Bravais Lattice Table" showing different suggested cells and their penalties. Appropriate crystal system can be selected by clicking on the "diamond" box on the left side of the "Bravais Lattice" name. Then, clicking "Apply" or "Apply and Close" sends this information to the HKL program. The later option, always suggested, closes the "Bravais Lattice Table" window. During this period, the "xdisplayf" window will also updated and show changes representing the indexing process. The "Bravais Lattice Table" window below shows the selection of a "primitive hexagonal system". Indexing of difficult crystals may require using reflections from multiple frames; consult the manual. 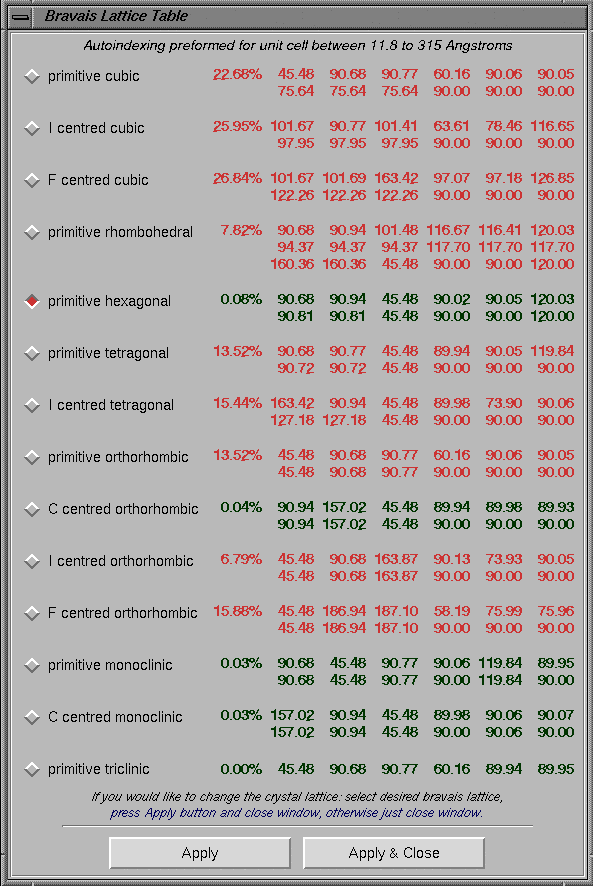
The HKL2000 program always select the lowest symmetry space group for a particular system during "Bravais Lattice" selection process (P2 for Monoclinic, P222 for Orthorhombic, P3 for hexagonal etc.). The correct space group can be selected during the Scaling process. The refinement of the cell can be started by clicking the "Refine" button. The parameters for refinement can be selected by either using specific buttons (Fit Basic or Fit All) or by activating the required parameters (Rot X, X Beam, etc.) by clicking on the box on the left side of the parameter. The spot size can be adjusted using the "Integration Box" window on the lower left corner of the above figure. Clicking on the "More Options" button brings additional parameters for advanced spot adjustments as shown below. Clicking on the "Simpler Options" window brings the previous simple view of the "Integration Box" window. (Note: Use the "xdisplayf" window to monitor the effect of box changes and other refinement specific parameter changes on the refinement.) 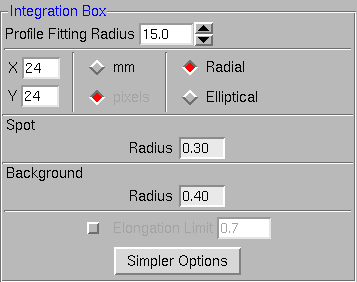
The refinement results for a well behaved crystal are shown below. 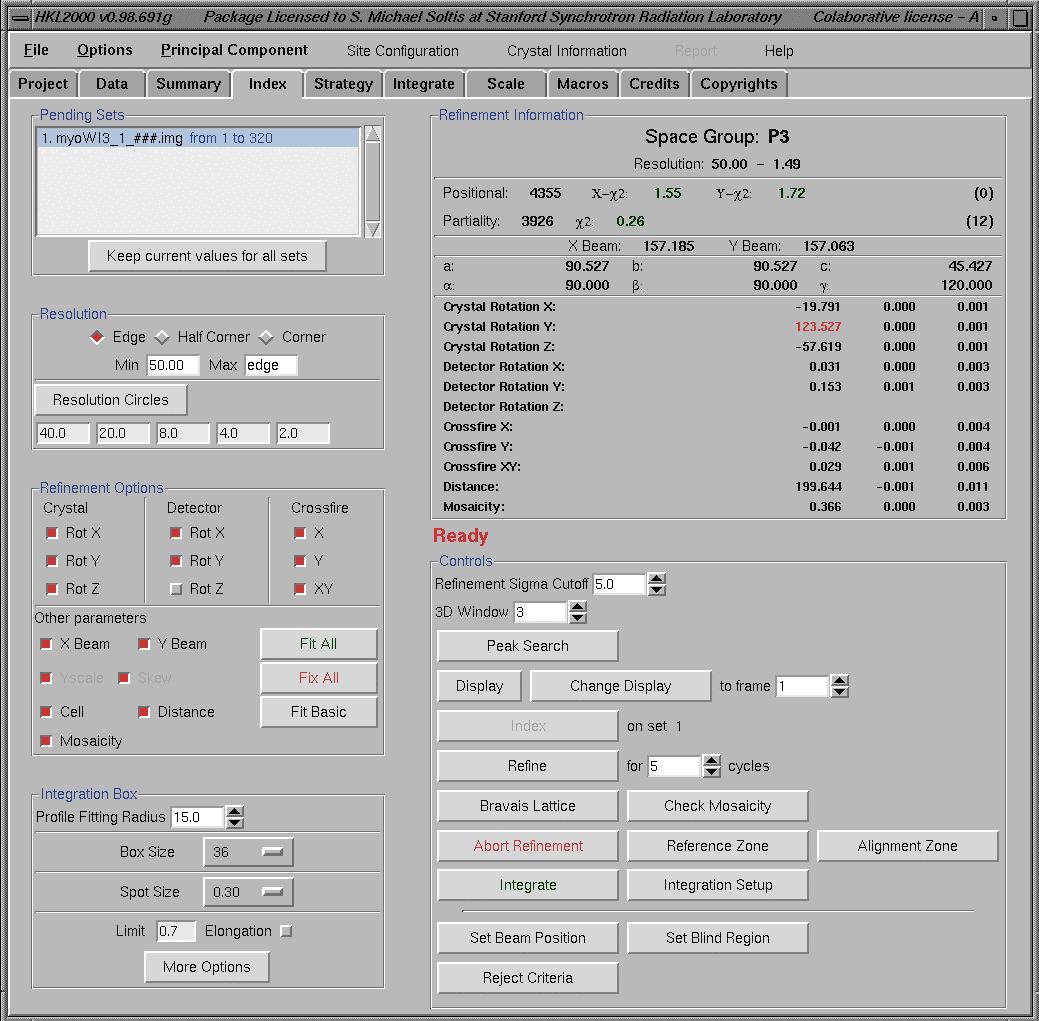
Setting Beam Position during index/refinementIt is also possible to define the beam position during indexing/refinement stage by selecting the Set Beam Position button from the bottom right part of the "Index/Refinement" window. (Note: The "Set Beam Position" box is active only when an image is displayed with the "Display" option.) This will bring up a small "Beam Position" window with detailed informations on how to select the beam position by clicking on a displayed image. The new beam position information can be either applied to the current refinement by selecting "Apply Current" or it can be entered into the Site Configuration window by selecting "Update Site". (Note: It is not possible to select the beam position from a zoomed window.) If the new beam position information was saved using the "Update Site" button, it is now possible to try indexing with the newly selected beam position after aborting the current refinement using "Abort Refinement" button. Please note that the indexing procedure always takes the "beam position" values from the "Site Configuration" file. Therefore, an indexing attempt without saving the newly selected beam position into the "Site Configuration" window will use the old beam position values. Setting Blind RegionA user defined blind region on the detector can be set by selecting the Set Blind Region box from the bottom right part of the "Index/Refinement" window. This will bring a small "Blind Region Coordinates" window with necessary information on how to define the blind region. Please note that this box will become active only when an image is displayed with the "Display" option. Selections are not possible on a zoomed window. Indexing with detector offsetSSRL beamlines use a simple horizontal or vertical translation to offset the detector. This offset information can be given to the program by entering the new beam center in the "Site Configuration" window. Please consult the User's Guide for help to determine the beam center. Strategy and SimulationAfter successful indexing of the images, the "Strategy or Simulation" programs can be accessed by selecting the "Strategy/Simulation" box on the top menu bar from any HKL window. The Strategy and Simulation programs provide information about the data completion and spot overlap, respectively. The "Strategy/Simulation" window below shows the results for a particular data set. 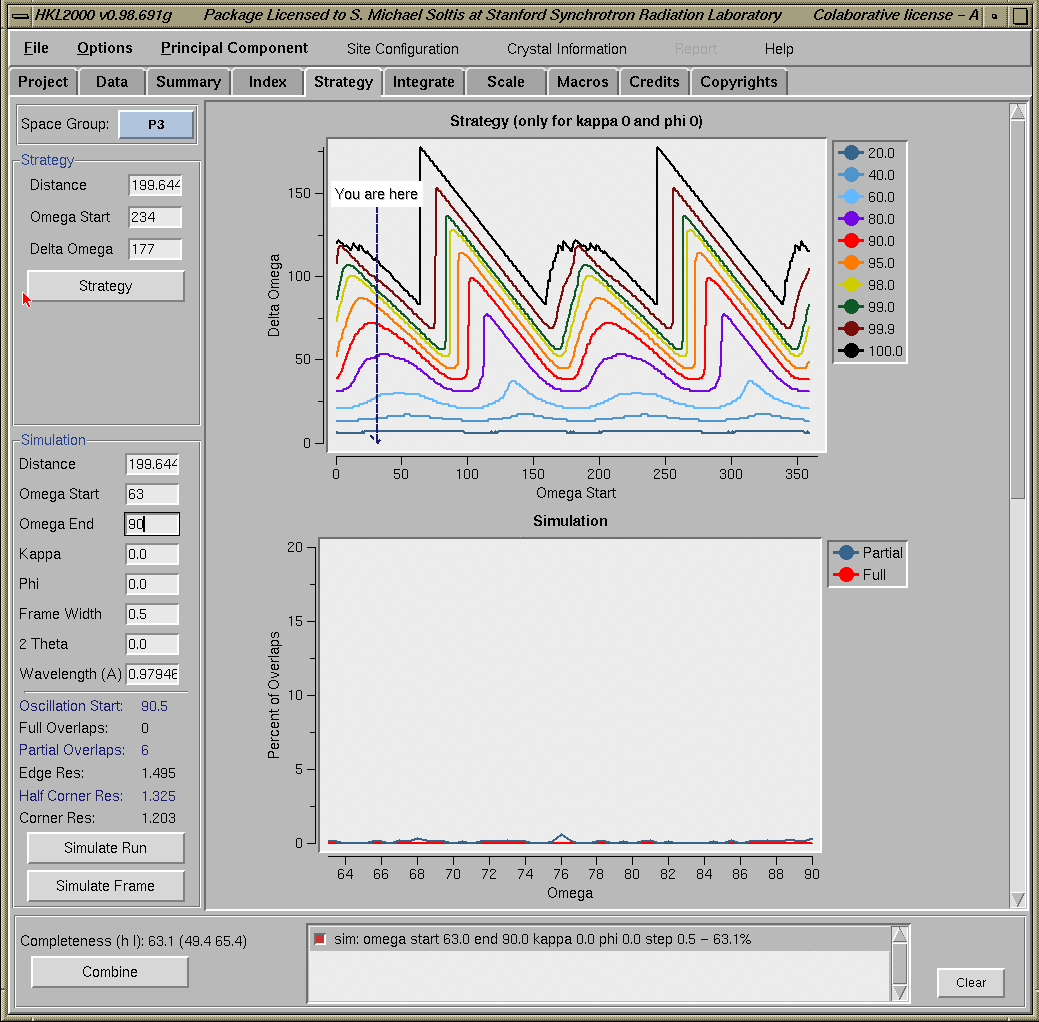
To obtain the optimal oscillation start, move the cursor to the first dip in the graph (black curve); the Y-Axis value at this position indicates the minimum oscillation range to achieve 100% completeness. The X-Axis value indicate the starting angle. If the number of overlaps is high (>5%), you may decrease the Frame Width or increase the Distance and rerun the Simulation until you are satisfied with the results. Note that the strategy results are independent on whether the sample actually rotates around omega or phi (the latter is usually the case at SSRL).IntegrationIntegration process can be started by clicking the "Integrate Sets" button from the "HKL Main" window. However, it is always recommended to go the "Integration" window (figure below) first, either by clicking on the "Integration Setup" button from the "HKL Main" window or by selecting the "Integration" menu from the menu bar on the top. This window allows to make changes in the integration script (by clicking on the "Special Integration" box) or to account for "Crystal Slippage" (by clicking on the "Crystal Slippage" box). Clicking the "Integrate Sets" button starts the integration process. 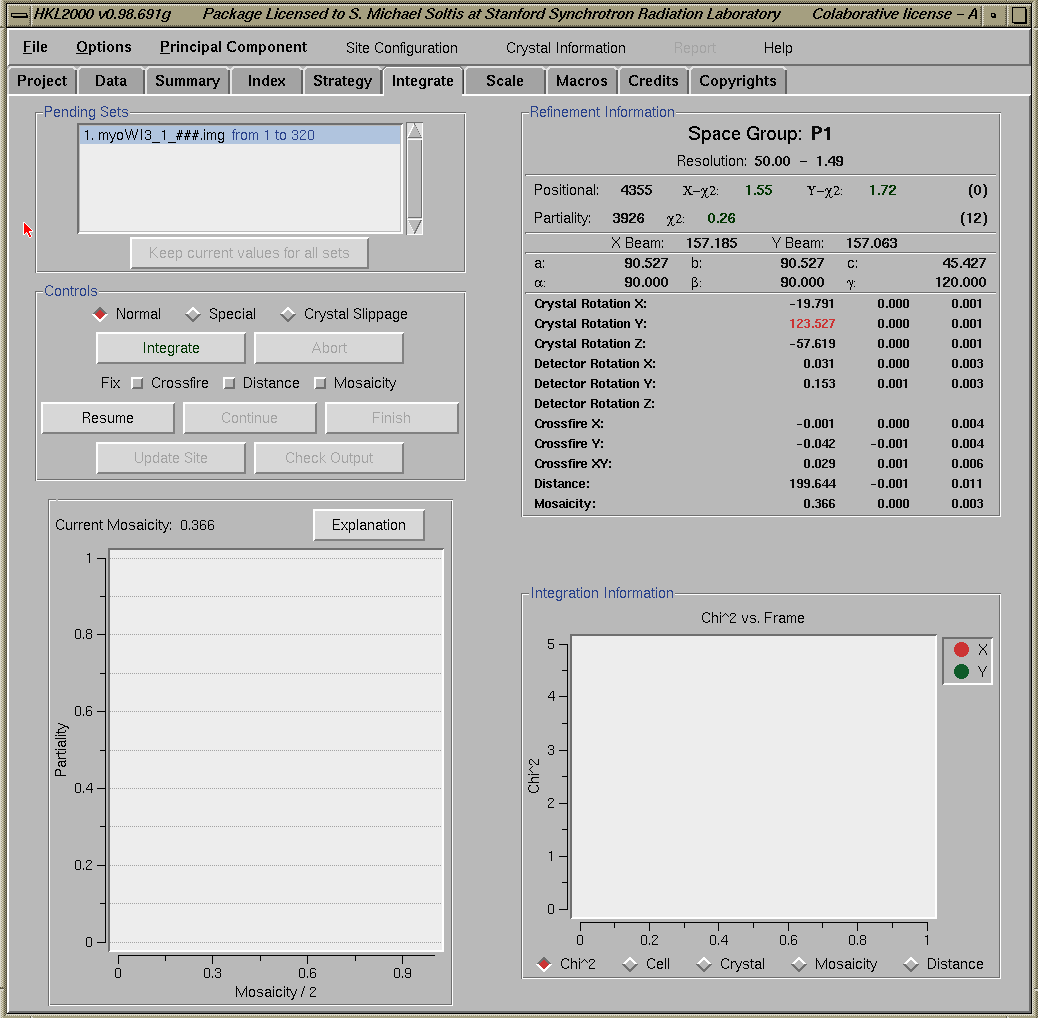
During integration, HKL2000 shows the progress of refinement with two graphs on the lower portion of the "Integration" window. While one of the graphs monitor crystal mosaicity for each set of frame (graph on the left side below), the second graph can display Chi square, Cell, Crystal, mosaicity, or Distance variations during the integration process (right side graph below shows Chi square variations). Clicking on each of these parameters at the bottom portion of the graph displays their respective graphs. The "xdisplayf" window also gets updated during the integration process. Information on how to adjust various parameters to obtain good agreement is available at the HKL2000 site
ScalingThe Scaling program can be selected by clicking on the "Scaling" box from the top menu of any "HKL window". A general view of the "Scaling window" before running scaling program is shown below. The upper half of the "Scaling window" can show 7 graphs (Scale and B vs frame, Completeness vs frame, etc.) at a time. These graphs can be viewed by moving the sliding bar on the right side of this window. 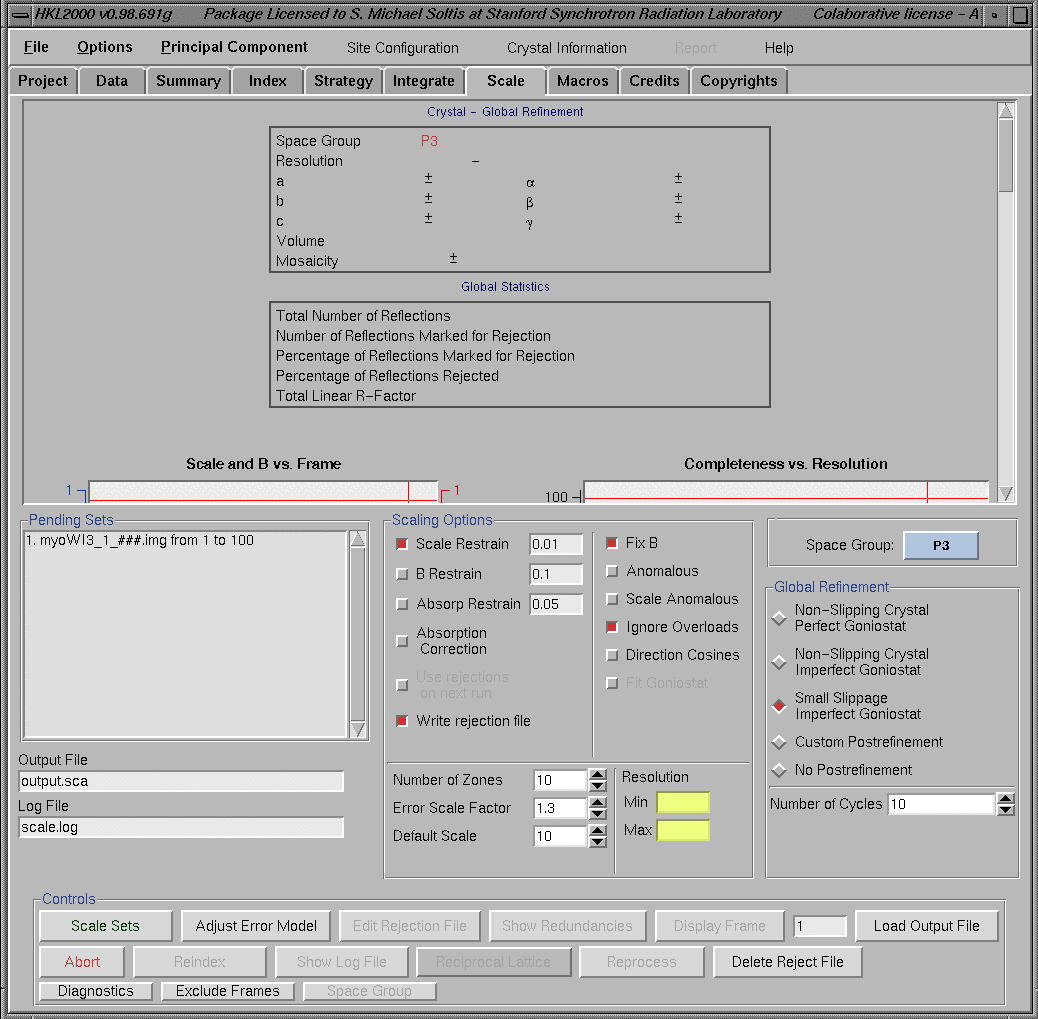
The correct space group can now be selected by choosing the "Space Group" selection button and navigating through the space group list. "Scaling window" also allows access to various parameters (number of zones, Error scale factor, etc.) and various scaling options (B restrain, Anomalous, etc.) at the lower section of the window. These options are fully documented in the HKL manual. Select the button on the left side of the "Use rejections on next run" to reject the bad reflections from a previous scaling run. Similarly select "Small Slippage Imperfect Goniostat" to activate postrefinement. After the initial round of scaling we can adjust the error model, edit rejection file, check log file, etc. from the "Controls" section on the bottom part of the "Scale" window. 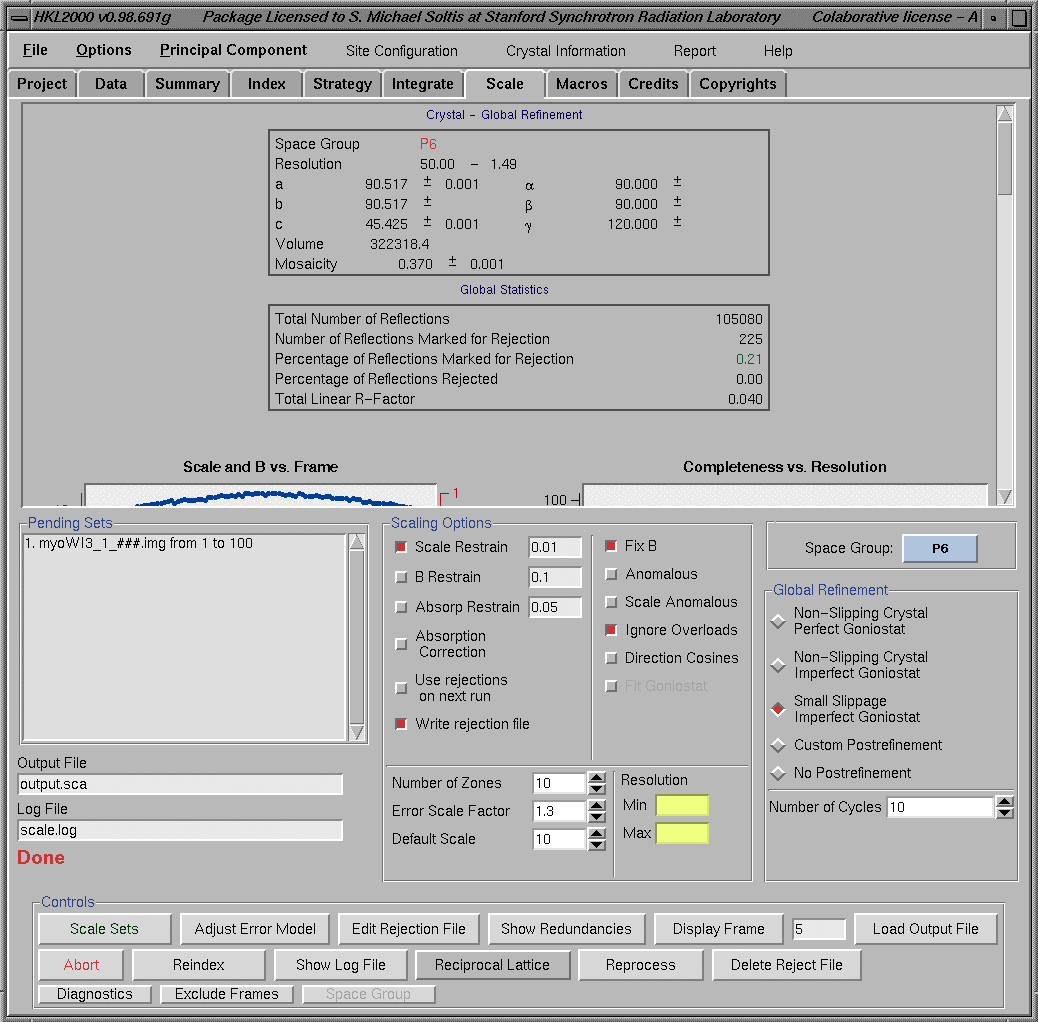
Selection of the Custom Postrefinement box from the "Global Refinement" menu opens a new section that allows selective refinement of various parameters during scaling. Running MacrosHKL2000 also has an option to execute "user created" macros during any of the data processing steps. The "Macros window" can be accessed by selecting "Macros" from one of the "HKL windows". The "Macros window" below shows the incorporation of "no merge original index" flag during scaling. To include a command into the macros, type the command on the line left to the "Add Macro" box and click on the "Add Macro" box. This will include that command on the macro area below. If necessary, this line can be removed by selecting it with a mouse button and clicking on the "Remove Macro" box. 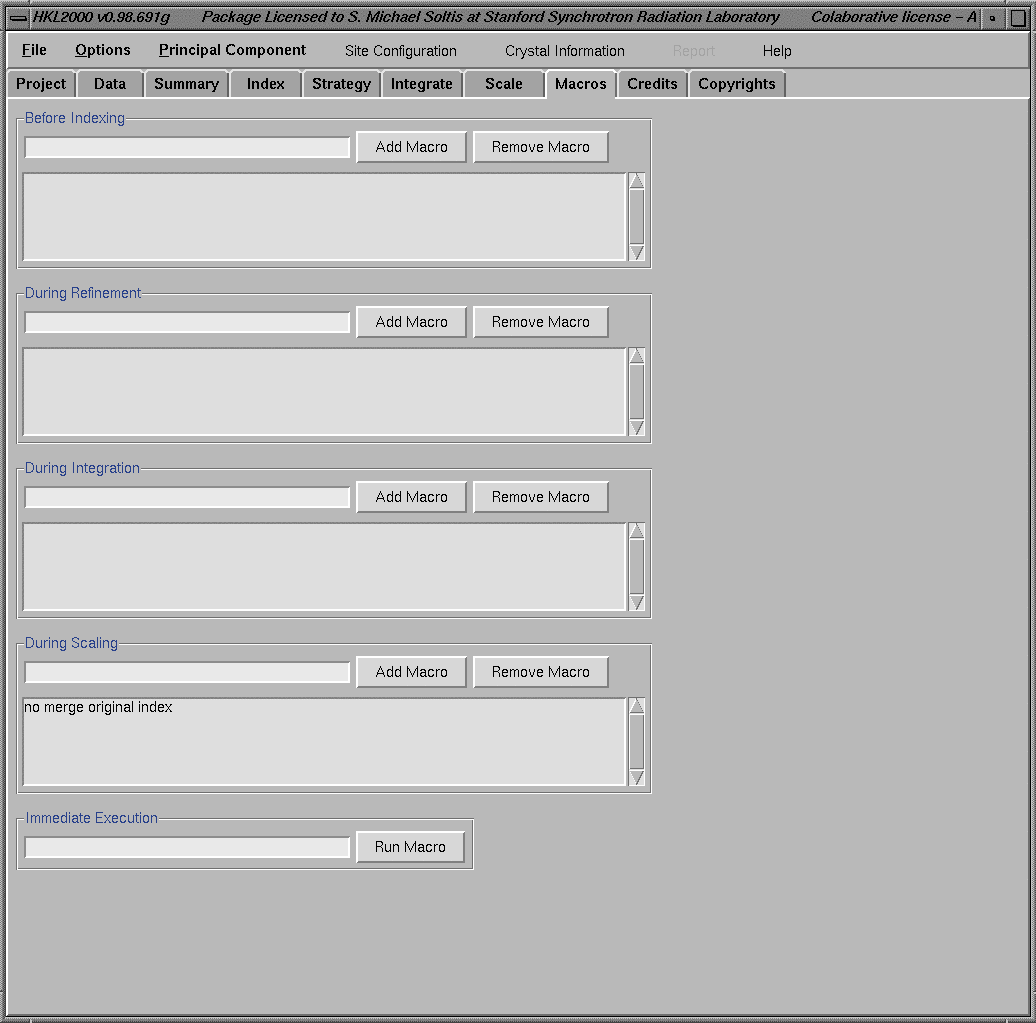
Bugs and ProblemsThis is a list of noticed problems with the latest version of HKL2000 (v0.98.691h):
Other HKL2000 Sites
|
||||
|
|
||||
| Technical questions: Webmaster
Content
questions: Irimpan Mathews |
||||
| Last modified:Wednesday, 06-Mar-2024 18:20:32 PST. | ||||